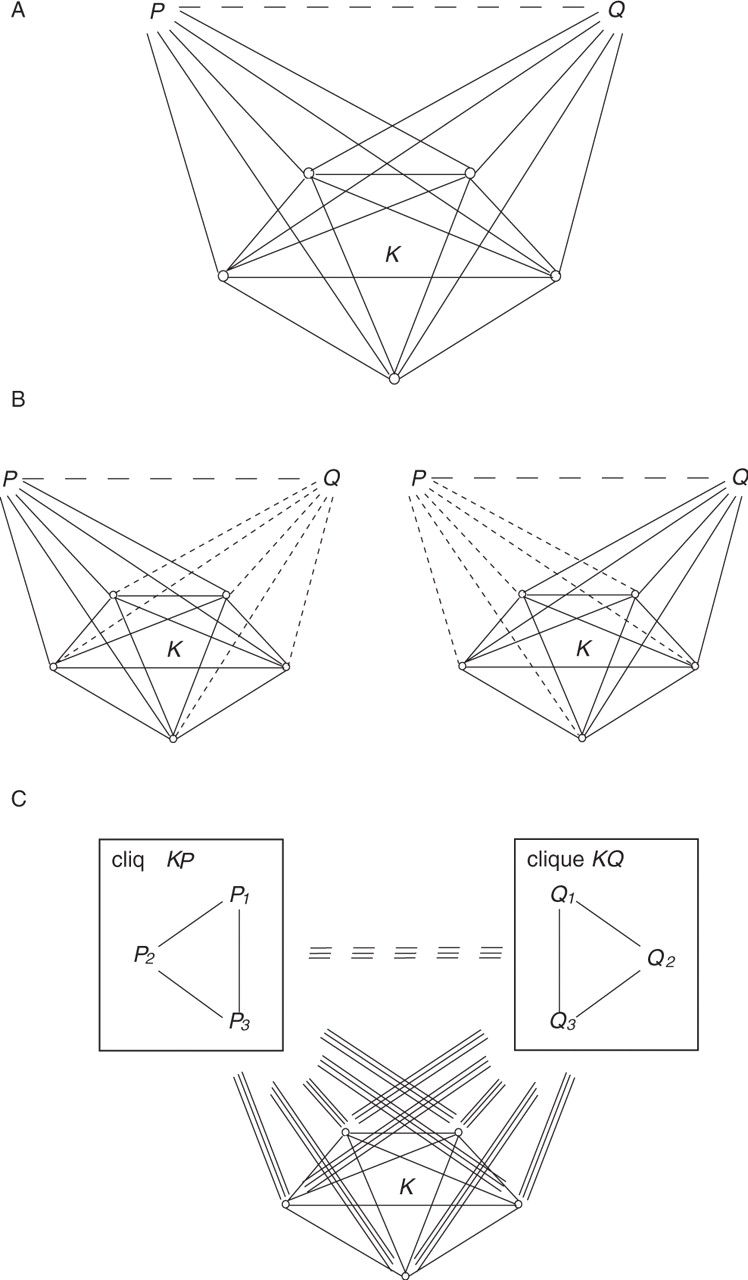
Schematic
illustrations of a defective clique and how the concept evolved. (A)
A defective clique in a protein interaction network. KP
and KQ
are both (k +
1)-cliques,
with k
overlapping vertices (i.e. clique K).
The dashed edge between proteins P
and Q
corresponds to a predicted interaction. KPQ
is a defective clique with a missing edge PQ. (B)
The decomposition of the defective clique (KPQ)
into the union of two overlapping cliques (KP
and KQ).
(C)
Generalized defective cliques. In general, a defective clique consists
of two cliques: K  KP
and K
KP
and K  KQ.
There are two parameters to determine a defective clique: k,
the size of the overlapping subclique (i.e. K);
l,
the size of the non-overlapping subcliques (i.e. KP
KQ.
There are two parameters to determine a defective clique: k,
the size of the overlapping subclique (i.e. K);
l,
the size of the non-overlapping subcliques (i.e. KP  KQ).
In the defective clique K
KQ).
In the defective clique K  KP
KP  KQ,
the dashed edges between subcliques KP
and KQ correspond to predicted interactions. [from H. Yu, A. Paccanaro, V. Trifonov, M.
Gerstein (2006). Predicting
interactions in protein networks
by completing defective cliques.
Bioinformatics 2006 Apr 1;22(7):823-9]
KQ,
the dashed edges between subcliques KP
and KQ correspond to predicted interactions. [from H. Yu, A. Paccanaro, V. Trifonov, M.
Gerstein (2006). Predicting
interactions in protein networks
by completing defective cliques.
Bioinformatics 2006 Apr 1;22(7):823-9]

 KP
and K
KP
and K  KQ.
There are two parameters to determine a defective clique: k,
the size of the overlapping subclique (i.e. K);
l,
the size of the non-overlapping subcliques (i.e. KP
KQ.
There are two parameters to determine a defective clique: k,
the size of the overlapping subclique (i.e. K);
l,
the size of the non-overlapping subcliques (i.e. KP  KQ).
In the defective clique K
KQ).
In the defective clique K  KP
KP  KQ,
the dashed edges between subcliques KP
and KQ correspond to predicted interactions. [from H. Yu, A. Paccanaro, V. Trifonov, M.
Gerstein (2006). Predicting
interactions in protein networks
by completing defective cliques.
Bioinformatics 2006 Apr 1;22(7):823-9]
KQ,
the dashed edges between subcliques KP
and KQ correspond to predicted interactions. [from H. Yu, A. Paccanaro, V. Trifonov, M.
Gerstein (2006). Predicting
interactions in protein networks
by completing defective cliques.
Bioinformatics 2006 Apr 1;22(7):823-9]